Mart van der Lugt
MSc student Drug Discovery / Data Science and AI Technology & Computer Scientist at Haaglanden Medical Center
Latest News
–
Presented my research on Pyrite at the NWO CHAINS conference.–
Completed my BSc in Pharmaceutical Sciences through the creation of Pyrite for my final thesis.–
Won both the audience and jury prize for best poster-presentation of the Molecular Toxicology course of the Pharmaceutical Sciences bachelor at VU Amsterdam.
Projects
Work Experience
Computer Scientist at Haaglanden Medisch Centrum
March 2022 - Present
- Maintaining and developing software in a clinical context within a small team.
- Statistical analysis and validation of patient and algorithm data.
- Organising and executing several R&D projects advancing patient care.
Teaching and Research Assistant at Vrije Universiteit Amsterdam
September 2024 - Present
- Teaching tutorials and computer practicals for the second-year Statistics course for several bachelor programs of the Faculty of Science.
- Assisting in practicals for the second-year Molecular Modeling course for Pharmaceutical Sciences.
- Coaching Pharmaceutical Sciences and Science Business and Innovation students during four-week projects.
- Developing websites for several research groups.
- Assisting BSc students with research methods during their thesis project in computational drug discovery.
Commissioner of External Affairs at VCSVU
June 2023 - June 2024
- In charge of all external contacts, educational activities and sponsoring for the study association for Pharmaceutical Sciences, VCSVU.
- Organised several successful educational events.
- Independently doubled contribution from sponsors.
Team Manager at ALDI Nederland
June 2018 - March 2022
- Managing a medium-sized team during peak hours in the weekend.
Publications
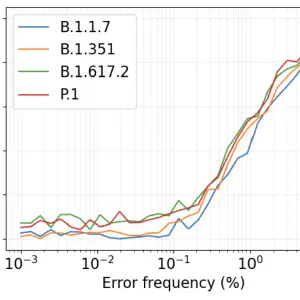
Baaijens, J. A.; Zulli, A.; Ott, I. M.; Nika, I.; van der Lugt, M. J.; Petrone, M. E.; Alpert, T.; Fauver, J. R.; Kalinich, C. C.; Vogels, C. B.; others. Lineage Abundance Estimation for SARS-CoV-2 in Wastewater Using Transcriptome Quantification Techniques. Genome biology 2022, 23 (1), 236. https://doi.org/10.1186/s13059-022-02805-9.